Functional Maturation of hiPSC Cardiomyocytes
hiPSC-derived cardiomyocytes closely resemble neonatal cells, limiting their predictive power for drug safety, toxicity, and disease modeling. Long-term electrical pacing overcomes this limitation by driving functional maturation in a consistent and scalable way.
Using the xCELLigence CardioECR system, hiPSC-CMs are reliably paced via planar impedance electrodes, resulting in improved gene and protein expression, enhanced contractility, and more adult-like electrophysiology. The outcome is a more predictive in vitro cardiac model for drug safety testing, discovery, and disease research.
Application Note:
Functional analysis of cardiac safety and toxicity assessment

Improved maturation via long-term electrical pacing.
More physiologically relevant responses.
Easy integration into high-throughput workflows.
Compatible with MEA, contractility, calcium, HCS, voltage dye assays, and plate readers.
Improved Gene Expression by Electrical Pacing
The expression of key maturation gene markers in non-paced (CTRL) and paced cardiomyocytes was determined by RT-PCR. The human fetal heart (FH) and adult left ventricle (ALV) were included as benchmarks of known level of maturation. The amount of RNA was normalized to the expression of GAPDH. Gene expression level was presented as a fold change to the human fetal heart. The data are presented as mean± SD, N=4. (* P< 0.05).
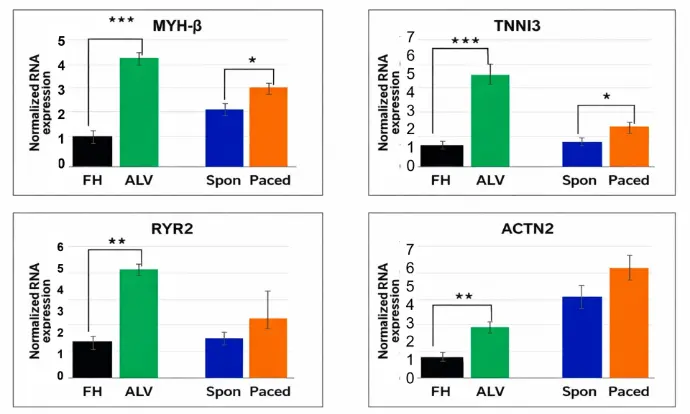
Improved Protein Expression by Electrical Pacing
Protein expression level of sarcomeric α-actinin, myosin heavy chain, cTNT, SERCA2a, and PLN in paced and non-paced cells were determined by Western blot analysis, which was quantified using protein to GAPDH ratio and further normalized to the non-paced cardiomyocytes. The data are presented as mean ± SD, N=4. (* P< 0.05)
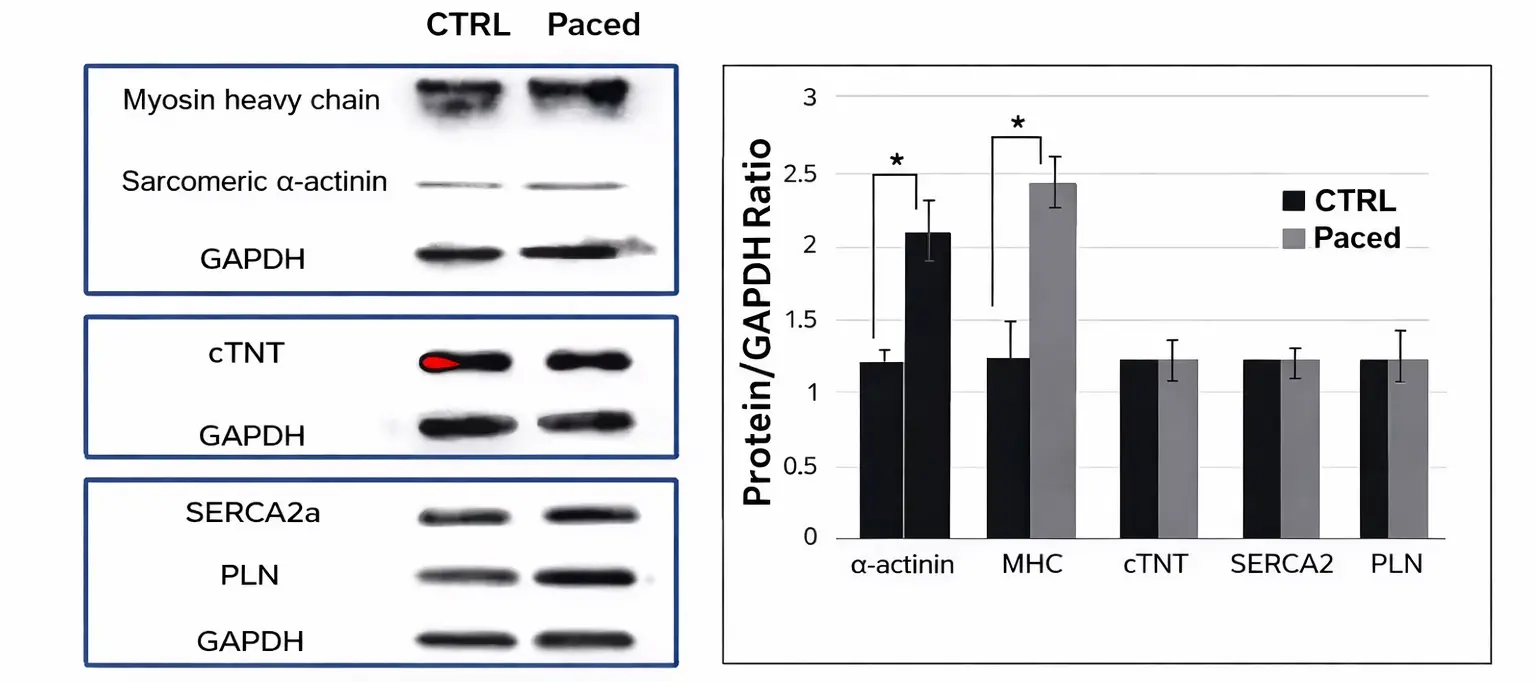
Improved Contractile Response by Electrical Pacing
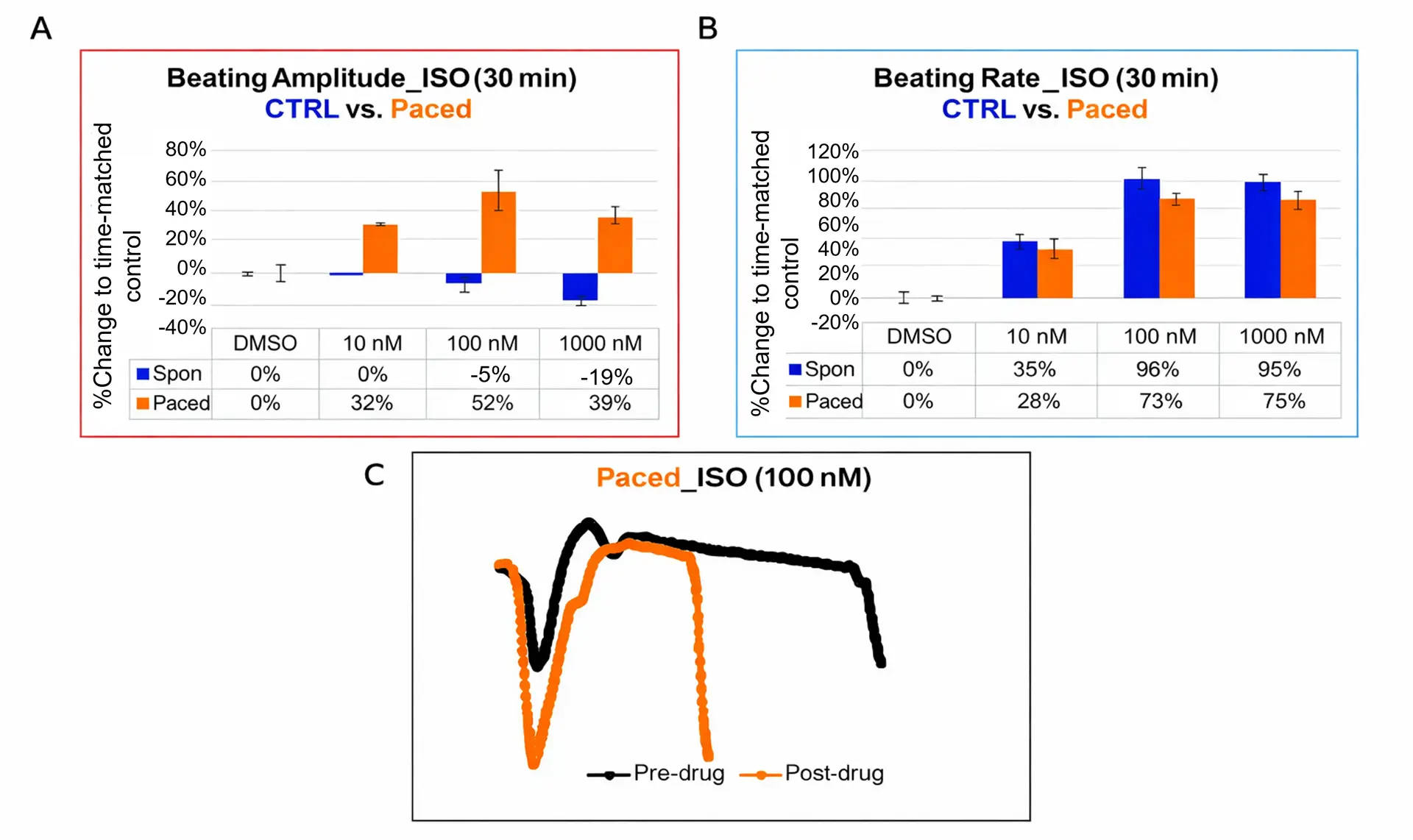
CTRL and paced cardiomyocytes were treated with isoproterenol (ISO; 10, 100, 1000 nM). Percentage changes in beating amplitude (a) and rate (b), normalized to time-matched DMSO controls, were measured 30 minutes post-treatment (CTRL: blue; paced: orange; mean ± SD, N = 3).
Impedance (IMP) waveforms were recorded before and after drug addition, averaged over 30-second intervals (c). Representative averaged waveforms before (black) and after ISO treatment in paced cells (orange) are shown (d).
Human iPSC-derived Cardiomyocytes
Human iPSC-CMs possess an inherent negative impedance amplitude-frequency relationship that is reversed after electrical pacing.
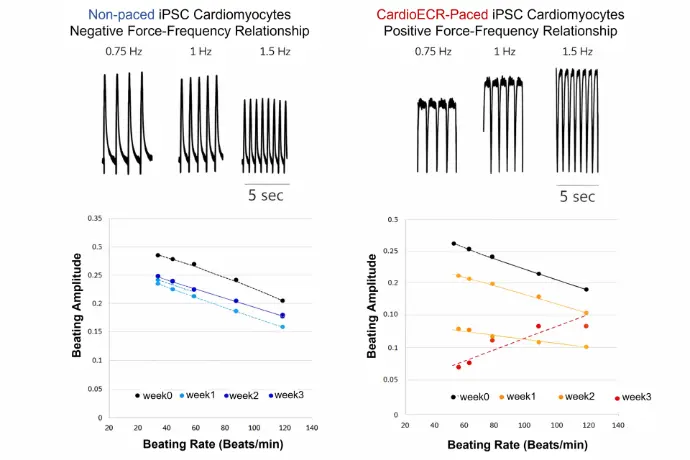
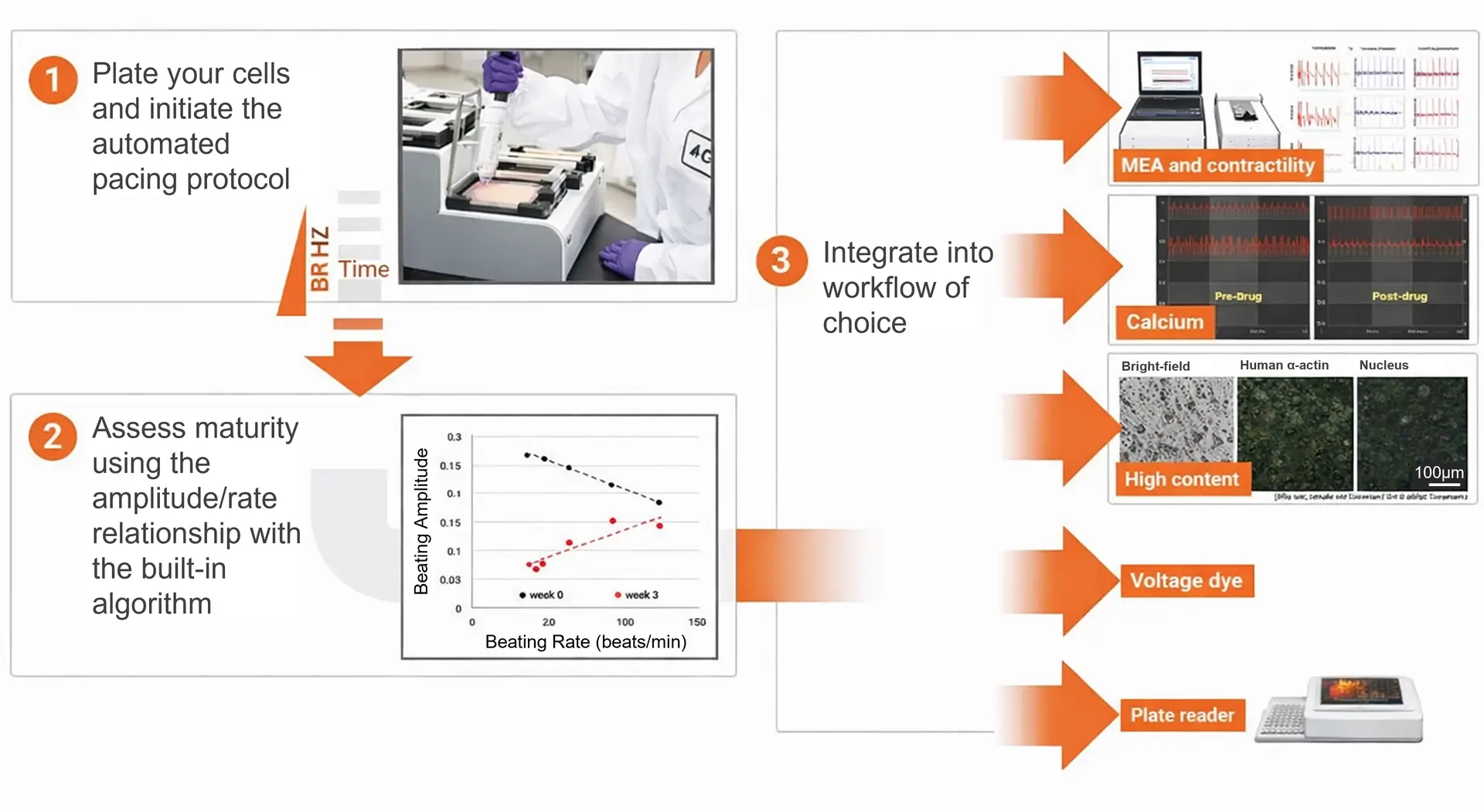
Easy, High Throughput and Versatile Workflow
The xCELLigence RTCA ePacer is adaptable and can easily integrate into your existing assay workflow. The view area on the E-Plate Cardio View 96 allows for compatibility with other optical assays. Designed to support the development of functionally mature cardiomyocytes, the ePacer easily fits into workflows such as multi-electrode array (MEA) measurements, contractility assays, calcium transient analysis, high-content screening (HCS), voltage-sensitive dye assays, and plate-based readers.
Application Note: Cardiomyocyte Maturation Using the xCELLigence RTCA ePacer
Related Application Notes
A Real-Time Impedance Analysis Instrument for Cardiac Safety and Toxicity Assessment
The Agilent xCELLigence RTCA CardioECR System
Assessment of Cardiomyocyte Disease Models Using the Agilent xCELLigence CardioECR
Functional Maturation of Human-Induced Pluripotent Stem Cell-Derived Cardiomyocytes
For Research Use Only. Not for use in diagnostic procedures.